IS Families/ISAzo13 family: Difference between revisions
Appearance
No edit summary |
No edit summary |
||
| Line 1: | Line 1: | ||
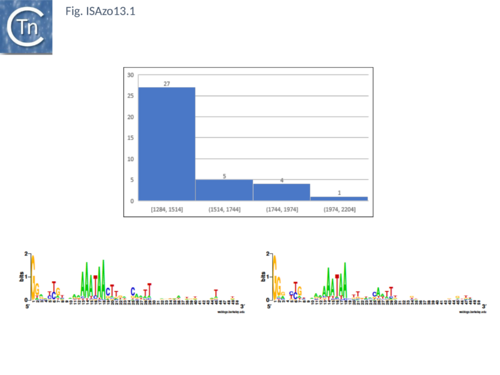
'''<big>T</big>'''his family, represented by 37 members in [https://isfinder.biotoul.fr/ ISfinder] emerged from the IS''NCY'' orphan group. It is based on both Tpase and IR sequence similarities [[:File:ISAzo13.1.png|(Fig.ISAzo13)]]. Insertion generates a 3bp AT-rich DR and the ends have a consensus GGa/g. Their Tpases are highly conserved with a probable DDE motif and an HTH motif at the N-terminus which could function as a DNA binding domain. Two members encode two orfs with a possible PRF (programmed transcriptional frameshifting) motif of 8 or 9 A while the other members encode a unique orf which includes a triple lysine at the equivalent position (Gourbeyre, unpublished). | '''<big>T</big>'''his family, represented by 37 members in [https://isfinder.biotoul.fr/ ISfinder] emerged from the IS''NCY'' orphan group. It is based on both Tpase and IR sequence similarities [[:File:ISAzo13.1.png|(Fig.ISAzo13)]]. Insertion generates a 3bp AT-rich DR and the ends have a consensus GGa/g. Their Tpases are highly conserved with a probable DDE motif and an HTH motif at the N-terminus which could function as a DNA binding domain. Two members encode two orfs with a possible PRF (programmed transcriptional frameshifting) motif of 8 or 9 A while the other members encode a unique orf which includes a triple lysine at the equivalent position (Gourbeyre, unpublished). | ||
[[Image:ISAzo13.1.png|thumb|center|500x500px|'''Fig. ISAzo13.'''|alt=]] | [[Image:ISAzo13.1.png|thumb|center|500x500px|'''Fig. ISAzo13.'''|alt=]] | ||
==Bibliography== | |||
</references> | |||
Revision as of 14:39, 1 June 2020
This family, represented by 37 members in ISfinder emerged from the ISNCY orphan group. It is based on both Tpase and IR sequence similarities (Fig.ISAzo13). Insertion generates a 3bp AT-rich DR and the ends have a consensus GGa/g. Their Tpases are highly conserved with a probable DDE motif and an HTH motif at the N-terminus which could function as a DNA binding domain. Two members encode two orfs with a possible PRF (programmed transcriptional frameshifting) motif of 8 or 9 A while the other members encode a unique orf which includes a triple lysine at the equivalent position (Gourbeyre, unpublished).
Bibliography
</references>